Tracing the signature of African-to-Neandertal gene flow
A new study of African genetic variation yields a more accurate picture of the genetic exchanges between ancient Africans and Neandertals 250,000 years ago.
A research study published last week by Daniel Harris and collaborators helps to answer an question that I've been wondering about: What's going on with the so-called “introgression deserts” in human genomes?
The study also provides more precise estimates for some patterns of mixture among ancestral human groups, including Neandertals. One finding confirms that later Neandertals inherited around 6% of their total genomic variation from African populations around 250,000 years ago. With new data from several African populations, the study clarifies the range of genetic input from Neandertals into African populations today.
The new research comes from an international collaboration led by the University of Pennsylvania human geneticist Sarah Tishkoff. Tishkoff's research group works with collaborators in many African nations toward a better understanding of genetic variation in African populations. Their work on the genetics of small-scale societies has been especially valuable, because these cultural groups are not represented in some of the massively funded genetics projects with explicitly biomedical aims, like the 1000 Genomes Project. In this study they included genomes from participants who belong to several indigenous groups including rainforest hunter-gatherers in Cameroon, and traditional hunter-gatherer and herding groups in southern Africa, Tanzania, and Ethiopia.
Neandertal ancestry in Africa
All populations in Africa have had historic connections with Eurasian populations, and gene flow from Eurasia during the last 40,000 years or more has brought some Neandertal ancestry into African peoples. In 2020, Lu Chen and colleagues from Joshua Akey's lab evaluated the Neandertal ancestry segments in the 1000 Genomes Project samples, which represent people from several large populations in Nigeria, Sierra Leone, Gambia, and Kenya. They found that around 0.6% of the genomes of these African populations had come from Neandertal ancestors. That's a bit less than a third of the Neandertal ancestry in most Eurasian populations, which is around 2%. Chen and coworkers found that most of the shared DNA from Neandertals could be explained by recent gene flow from Eurasian groups, and not a more ancient mixture with Neandertals.
But the 1000 Genomes Project participants don't represent all the variation of African peoples. Traditional small-scale societies have a varied pattern of ancestry, some of them with less historic mixture from Eurasian populations. Some of the groups included in the new research by Harris and coworkers had as much or more Neandertal ancestry as the 1000 Genomes Project samples studied by Chen and coworkers. These groups include some groups from Ethiopia and Tanzania, while others have less Neandertal ancestry. Groups in southern Africa and rainforest hunter-gatherers have the lowest Neandertal ancestry, ranging down to a level low enough that the samples cannot distinguish it from zero. As in the earlier research, these Neandertal ancestry segments basically track the amount of genetic ancestry that individuals have within the last 40,000 years from Eurasian source populations.
Six percent of Neandertal ancestry was African
Within the last ten years, research has shown that Neandertals received substantial gene flow from some popuations that were much more like African ancestors of today's people. Two of the most striking outcomes of this gene flow were the total replacement of early Neandertal mitochondrial and Y chromosome variation with new variants that originated in Africa. In 2020, Melissa Hubisz and coworkers from Joshua Akey’s research group quantified the early genetic input from Africans into Neandertals. They could show that around 3% of the Neandertal genome looked like segments from an African source and they estimated that their method could only detect around half of the ancient introgression, leading to an overall estimate around 6%.
In their new study, Harris and coworkers were able to leverage the greater representation of present-day African variation to make a more precise estimate of ancestral African-to-Neandertal gene flow. Their estimate of 6% confirms that Hubisz and collaborators had the answer close to correct.
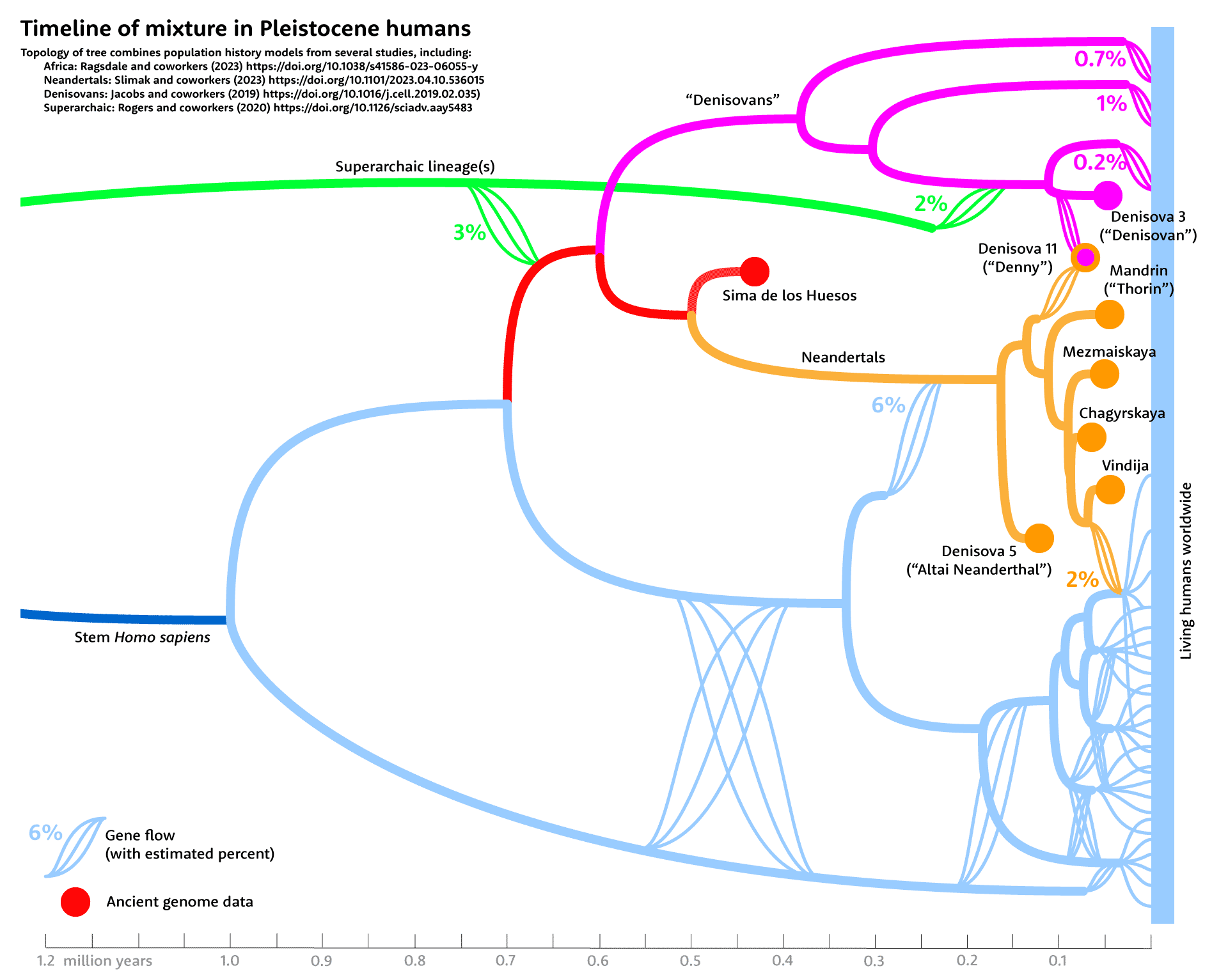
Both approaches agree that this African-to-Neandertal gene flow was underway in the time period around 250,000 years ago. This time range is also suggested by the ancient DNA data on mtDNA from early Neandertals. The data do not show whether the introgression from Africans happened in a rapid burst or many contacts over a prolonged time.
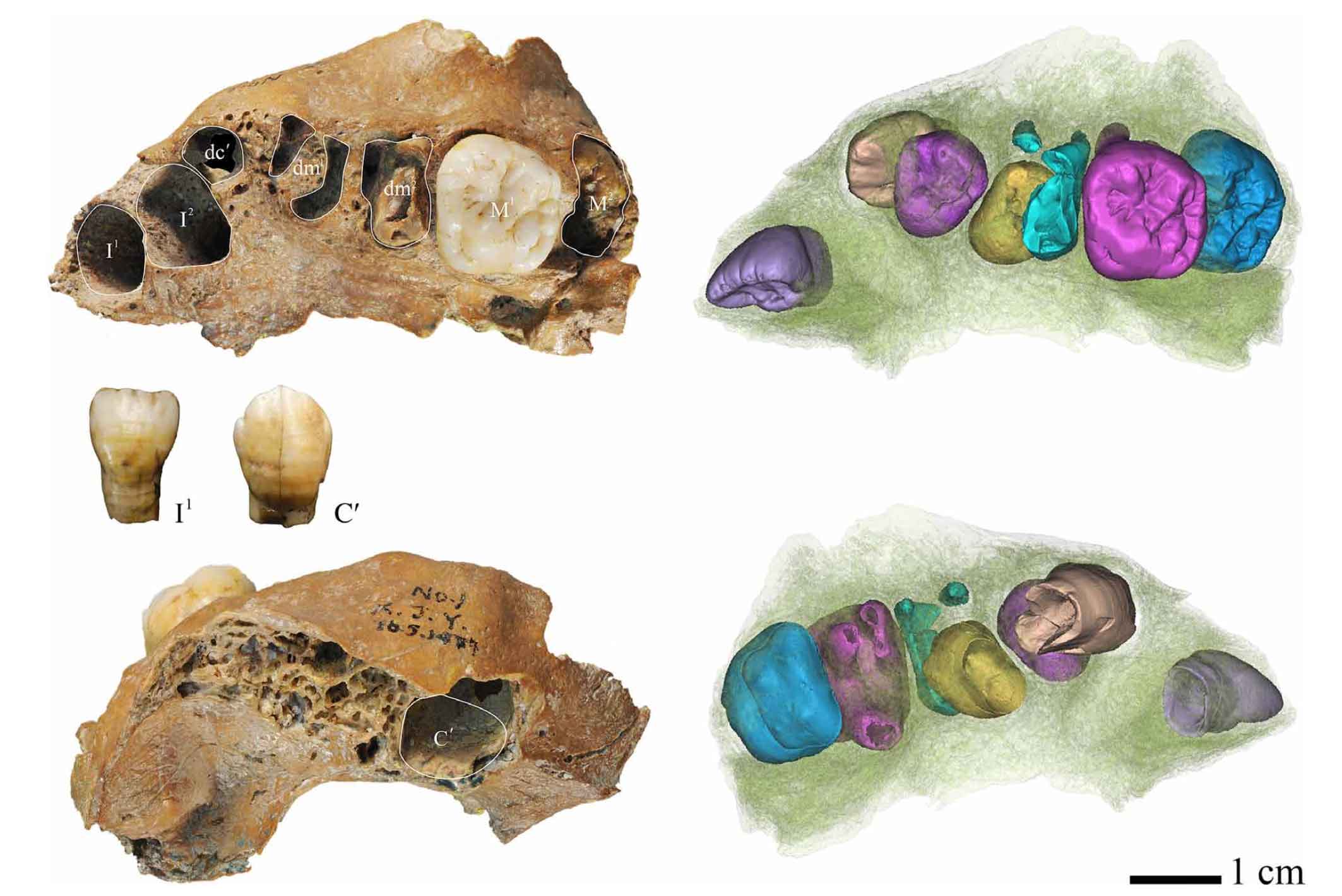
So far, those of us who study fossils don’t really know when and how this early mixture happened. We have a few fossils from Europe that seem like they might have more African than Neandertal input, including the Apidima 1 partial cranium from Greece which is more than 210,000 years old. To me the key region is southwest Asia, and our record there before 200,000 years ago is pretty spotty. The most interesting fossils may be some teeth from Qesem Cave, Israel, which have some Neandertal and some non-Neandertal aspects. But they’re just a few teeth. Nothing is really all that convincing in showing interaction of populations at this time period, and that’s just the nature of the fossil record.
One thing is for certain: This gene flow from an African source population shaped the evolution of Neandertals. The time period between 300,000 and 200,000 years ago was around halfway between their initial diversification from Denisovan ancestors and their last meeting with African groups. Every Neandertal sampled after this time had a mitochondrial genome derived from Africa; every individual with a Y chromosome was likewise part of an African clade. From central Asia to Iberia, the African genes spread everywhere.
But not all the African genes were beneficial. Harris and coworkers uncovered evidence that some of the African-to-Neandertal gene flow was deleterious.
What introgression deserts are
An introgression desert is a region of a chromosome that has an unusually low fraction of Neandertal or Denisovan derived sequence variants in a sample of living people, when compared to the average across the entire genome. The genetic ancestry that any living person has from these source populations is slight—around 2% for Neandertal ancestry—and this means that any one person's genome may include many regions of a million base pairs or more where this ancestry is absent. When we line up many people's genomes, these regions tend to be a random scatter. But not entirely. Looking at thousands of people, there are some parts of chromosomes where almost nobody has any Neandertal ancestry. These areas are the introgression deserts.
The most studied cases are on the X chromosome. The X has much less introgression than the autosomes and some extended regions of the X seem to have little or no Neandertal ancestry at all. I wrote about this earlier this year (“Explaining the X chromosome hole in Neandertal ancestry”). There are many other deserts across other chromosomes, shorter, and sometimes surrounded by regions with average or even higher-than-average Neandertal ancestry.
It's not yet entirely clear how much these Neandertal introgression deserts are also Denisovan introgression deserts. Ancestry from Denisova-like populations has been more challenging to study than Neandertal ancestry. Denisova-like genetic ancestry is most common today in Papua and nearby areas of island southeast Asia and Oceania, all underrepresented populations in human genetic research. Living populations that have much larger genome samples, mainly from mainland Eurasia, have only a tenth as much Denisovan ancestry. Today the best information about possible Denisovan introgression deserts comes from a sample of more than 27,000 genomes from Iceland, studied by Laurent Skov and collaborators in a 2020 paper. (I discussed these results here: “An enormous sample sheds light on the Denisovan ancestry of people in Iceland”.) Across these people's genomes, there is some overlap in chromosome regions that underrepresent Denisova-like ancestry and Neandertal ancestry. This overlap provides some basis for thinking that introgression from both ancient groups might follow similar dynamics in later populations, but more information about Denisovan ancestry will clearly be needed.
Of course it's also important not to lose sight of the fact that some Neandertal variants actually increased in frequency in modern populations: that is, introgression introduced some advantageous gene variants, along with many neutral ones. What is interesting about the introgression deserts is that explaining them may give some hints to the dynamics of mixing between ancestral groups, as well as their degree of compatibility after hundreds of thousands of years of evolution.
Explaining introgression deserts
Geneticists have proposed several ideas about the introgression deserts. The first idea, soon after the X chromosome Neandertal ancestry deficit was first identified, was that Neandertals and modern individuals may have faced incompatibilities when they mated. In this scenario, certain gene variants from Neandertals would have very low fertility in hybrid individuals, possibly even causing sterility in one sex. The phenomenon of hybrid incompatibility has been observed in many sister species of mammals. What such sister species tend to lack is the widespread evidence for gene flow that ancient DNA has demonstrated among Neandertals and their contemporaries.
Two more current hypotheses involve more subtle aspects of natural selection on introgressed genes, operating over more generations. One is genetic load, a phenomenon in which a population becomes more and more saddled with slightly deleterious genetic variants. What we know about Neandertal population structure suggests that they lived in small local populations with high inbreeding relative to most of today's human populations. With smaller population numbers, natural selection would have been less effective against slightly deleterious genetic variation in the Neandertal population. Over thousands of generations, the Neandertals may have accumulated thousands of slightly deleterious gene variants. The early modern people who inherited these slightly deleterious variants belonged to a growing population, in which purifying selection became more effective. Children with the slightly deleterious Neandertal variants would have been a little less likely to survive and have children of their own. Over hundreds of generations, parts of the genome with the most deleterious variants ended up with much less Neandertal ancestry, some with none at all.
Still, genetic load would not have been unique to Neandertals. The ancestral African population may have been a bit larger than the overall Neandertal population, but it also was organized into regional groups with relatively high inbreeding compared to recent times. The founder population that left Africa and encountered Neandertals was especially small and constrained in variation. In some parts of the genome, we might expect to see Neandertal genes actually increase in the subsequent population because of the founder population's genetic load.
The alternative hypothesis is genetic epistasis. No gene works in isolation, all genes must work in a context determined by many other genes across the entire genome. If a gene is taken out of one population and placed into another, it may interact in unexpected ways with its new genetic background. Introgressed genes from a population that first separated more than 20,000 generations in the past might sometimes have deleterious effects, resulting in their loss over time.
The two hypotheses are not mutually exclusive. Some introgressed segments might be slightly deleterious in modern populations because they were already slightly deleterious in Neandertals—that's the genetic load hypothesis. Others might be slightly deleterious in modern populations because they don't work the same as they did in Neandertals—that's the epistasis hypothesis.
One way to test the difference is to look at gene flow in the opposite direction: What happened to genes that originated in African populations and introgressed into Neandertals?
Harris and collaborators did exactly that. They considered areas of chromosomes where Neandertal introgression is low or absent in today's people, and they looked at the same regions in Neandertals. What they found is that the 6% African-to-Neandertal introgression is reduced by around half in these introgression deserts. They interpret this observation as evidence of epistasis. Neandertal genes in these regions do not work as well in today's genetic background; early African genes in the same regions did not work as well in the Neandertal genetic background.
On the other hand, the African source population for this ancient introgression was also likely to have been small in numbers. To me this suggests that genetic load may have mattered reciprocally in both ancient populations. The fact that the same regions seem to have had more slightly deleterious introgressed variants in both directions may tell us more about the strength of purifying selection in these regions, rather than the mechanism by which variants became deleterious.
Bottom line
The results in this paper are helping clarify the pattern of selection on ancient introgression. It's fascinating that Neandertals around 250,000 years ago experienced a pattern of purifying selection that we also recognize in more recent humans within the last 100,000 years. The magnitude of selection seems to be less—only half as much in these regions of the genome—and that may simply reflect the lower genetic distance between Neandertals and their early African contemporaries in the later Middle Pleistocene.
What about the idea of some genetic incompatibility? If these introgression deserts do indeed result from epistasis between genes in each region and the wider genetic background, they may be building blocks for speciation between these ancient human groups. At the moment, comparisons across the whole genome are just too coarse to look at how gene function relates to the genetic background. The next step should be to highlight the regions that are giving consistent outcomes in both today’s populations and ancient genomes, because those consistent regions are likely to actually have genes that matter to adaptation in both populations.
Notes: Several groups of researchers have investigated Neandertal introgression and its possible genetic costs in their descendants. I've written about this topic a number of times, and covered some of the concepts before. In the references below, I've included works by authors including Kelley Harris and Rasmus Nielsen, and Ivan Juric and coworkers, who explored the idea of genetic load in small ancestral Neandertal groups. I've also included references that I relied on for estimates of gene flow and ancestral relationships in the network diagram above.


References
Chen, L., Wolf, A. B., Fu, W., Li, L., & Akey, J. M. (2020). Identifying and Interpreting Apparent Neanderthal Ancestry in African Individuals. Cell, 180(4), 677-687.e16. https://doi.org/10.1016/j.cell.2020.01.012
Harris, D. N., Platt, A., Hansen, M. E. B., Fan, S., McQuillan, M. A., Nyambo, T., Mpoloka, S. W., Mokone, G. G., Belay, G., Fokunang, C., Njamnshi, A. K., & Tishkoff, S. A. (2023). Diverse African genomes reveal selection on ancient modern human introgressions in Neanderthals. Current Biology. https://doi.org/10.1016/j.cub.2023.09.066
Harris, K., & Nielsen, R. (2016). The Genetic Cost of Neanderthal Introgression. Genetics, 203(2), 881–891. https://doi.org/10.1534/genetics.116.186890
Hubisz, M. J., Williams, A. L., & Siepel, A. (2020). Mapping gene flow between ancient hominins through demography-aware inference of the ancestral recombination graph. PLOS Genetics, 16(8), e1008895. https://doi.org/10.1371/journal.pgen.1008895
Jacobs, G. S., Hudjashov, G., Saag, L., Kusuma, P., Darusallam, C. C., Lawson, D. J., Mondal, M., Pagani, L., Ricaut, F.-X., Stoneking, M., Metspalu, M., Sudoyo, H., Lansing, J. S., & Cox, M. P. (2019). Multiple Deeply Divergent Denisovan Ancestries in Papuans. Cell, 177(4), 1010-1021.e32. https://doi.org/10.1016/j.cell.2019.02.035
Juric, I., Aeschbacher, S., & Coop, G. (2016). The Strength of Selection against Neanderthal Introgression. PLOS Genetics, 12(11), e1006340. https://doi.org/10.1371/journal.pgen.1006340
Rogers, A. R., Harris, N. S., & Achenbach, A. A. (2020). Neanderthal-Denisovan ancestors interbred with a distantly related hominin. Science Advances, 6(8), eaay5483. https://doi.org/10.1126/sciadv.aay5483
Sankararaman, S., Mallick, S., Dannemann, M., Prüfer, K., Kelso, J., Pääbo, S., Patterson, N., & Reich, D. (2014). The genomic landscape of Neanderthal ancestry in present-day humans. Nature, 507(7492), Article 7492. https://doi.org/10.1038/nature12961
Slimak, L., Vimala, T., Seguin-Orlando, A., Metz, L., Zanolli, C., Joannes-Boyau, R., Frouin, M., Arnold, L. J., Demuro, M., Devièse, T., Comeskey, D., Buckley, M., Camus, H., Muth, X., Lewis, J. E., Bocherens, H., Yvorra, P., Tenailleau, C., Duployer, B., … Sikora, M. (2023). A late Neanderthal reveals genetic isolation in their populations before extinction (p. 2023.04.10.536015). bioRxiv. https://doi.org/10.1101/2023.04.10.536015
John Hawks Newsletter
Join the newsletter to receive the latest updates in your inbox.