Ghostbusters of human origins
Humans tend to mix and interact with each other. Geneticists are once again starting to take that seriously, changing their view of our origins.
Ghost populations are one of the biggest topics in human evolutionary research today. The term refers to ancient groups that no longer exist, but that left footprints of ancestry within the genomes of more recent people. Geneticists identify them by highlighting long, linked stretches of DNA that are unusually divergent and found only within a few populations. Sometimes ancient DNA turns out to match the genetic signature of a ghost population, confirming that it really existed. Without such proof, we may be left wondering if a putative ghost is just a figment of a mathematical model.
Ghosts are on my mind because of a new paper by Aaron Ragsdale and collaborators. In it, they try an older—but challenging to model—way of looking at the population history of Africa during the last million years. Where recent work leaned on ghost populations to explain current African diversity, Ragsdale and coworkers applied the common sense idea that past populations probably interacted a lot like recent ones have. They found this idea does a better job of explaining today's genetic variation, without ghost populations. Instead, ancestral Africans lived in a network of groups that have been interacting with each other for most of the last million years.
This way of looking at evolution is a throwback in some ways to work that preceded the revolution in ancient DNA. After 2010, the sequencing of the first Neandertal genome had far-reaching effects on population modeling, causing the ghost train to build up steam. But models with more substantial interactions among groups may resemble the way that ancient humans really behaved.
Ghosts in Africa
For the last decade, most reconstructions of African population history have been seeing ghosts. This got started as early as 2006, when Vincent Plagnol and Jeffrey Wall looked at West African and European population samples to consider how much of their genomes might have come from divergent ancestral groups. They estimated that 5% of European ancestry came from a very divergent group, which they speculated was Neandertals. This was later confirmed in the general sense by Neandertal ancient DNA evidence, which showed real but somewhat less introgression from this population in recent Europeans. Meanwhile, Plagnol and Wall also found that around 5% of West African ancestry likewise came from a divergent ancestral group, different from the first. They suggested that a parallel population of archaic humans had existed in Africa and mixed in with the other ancestors of modern people: a ghost population.
At the time, geneticists were busily adding new samples of single nucleotide polymorphism (SNP) data, mostly from research participants from the U.S., Europe, Japan, and China. Microarray SNP data became the building block of gene-phenotype association studies, which came to rely on linkage between SNP loci to narrow down small parts of the genome that make a difference to traits. But these SNPs had been identified mostly in people from the U.S. and Europe. Better understanding of African population history required whole genome sequences from African populations.
When human geneticists began to study whole genomes from Africa, they found patterns that seemed to reflect a longer period of evolutionary diversity than their previous data, limited to SNPs and mtDNA, had led them to expect. In 2012, Joseph Lachance and coworkers suggested that a ghost contribution could explain some aspects of the variation of whole genome sequences from African foraging groups. Later work followed, as geneticists investigated more populations. One of the strongest recent analyses, by Arun Durvasula and Sriram Sankararaman, modeled ghost introgression in West African ancestry populations, estimating between 2% and 19% of their ancestry came from unknown ancestral groups.
Ancient DNA from African sites made the picture more interesting. DNA survives longer in colder environments, which makes Africa a very challenging place for long-term DNA persistence. No one has yet recovered DNA from hominin material in Africa older than 20,000 years. Still, geneticists have succeeded in DNA retrieval from many later sites in Africa. The oldest have given some hints about the location and relationships of human groups before early herding and farming cultures began to grow and spread.
Carina Schlebusch and coworkers made news in 2017 with some of the first ancient DNA from southern Africa. The skeletal remains they studied, only a few thousand years old, were ancestors or relatives of living people of Khoesan language-speaking groups. Compared to these ancient relatives, genomes of today's Khoesan peoples show a few more genealogical connections to other populations in Africa and Eurasia. The closeness of today's people partially resulted from gene flow connecting peoples during the Holocene, from the spread of Bantu-speaking peoples and trade routes connecting peoples across the continent. Schlebusch and collaborators saw that the slightly greater genetic diversity of early Holocene people might imply a deeper evolutionary history. They suggested a model in which today's populations belong to a tree with its deepest branches starting around 300,000 years ago. At the same time, a group of paleoanthropologists led by Jean-Jacques Hublin provided new age estimates for early human fossils and Middle Stone Age artifacts from Jebel Irhoud, Morocco, placing them between 250,000 and 350,000 years old.
More ancient DNA work built from this timeline, showing a large role for ghost populations. Mark Lipson and collaborators in 2020 reported on ancient DNA from Shum Laka rockshelter, Cameroon. Again, these were skeletal remains of early Holocene people, their genomes showing connections to some of today's forest-living peoples of central Africa. As Lipson and coworkers built the tree connecting genomes from varied African populations, they found that they needed more deep branches to explain them. The resulting trees included “ghost archaics” and “ghost moderns”, unknown populations from two different nodes.

This kind of research recognizes that genomes of individuals from the same population have a shared history of genetic drift, and also may have ancestry from other populations by gene flow. The lengths of haplotypes shared by different individuals and the alleles that differentiate those haplotypes are outcomes of that history, and geneticists can fit a model of population branching, ancestral population sizes, and gene flow to those data. Long haplotypes with many alleles separating them from other haplotypes in the population are more likely to have been introduced by gene flow. When geneticists can't find these or similar haplotypes in other population samples, they assume that they came from some population that they haven't sampled. This is a ghost population.
Weakly-structured stems
A ghost population is one solution to a mathematical equation that relates genetic drift and gene flow when populations are strongly isolated and gene flow occurs in short bursts. There are other ways to look at genomes. If genetic exchanges were more continuous over time, the equation changes. What seems like a ghost may dissolve.
Ragsdale and colleagues began their work with the idea that long periods of isolation between groups in Africa just weren't very realistic when you look at how recent people behave. From the outside, it may appear that many small-scale human societies have been culturally isolated. Most have been faced with exploitation by colonizing population for hundreds of years, requiring high social solidarity for cultures and languages to persist. But cultural persistence is not demographic isolation. Small-scale human societies have high rates of intermarriage across cultural and linguistic divides. Yet most geneticists have relied on models that assume early members of our species lived in small groups that remained in hermetic isolation for tens of thousands of generations. Ragsdale and coworkers wondered whether a model with more gene flow might yield different results.

Gene flow does matter. A model with long, continuous gene flow among the most ancient human groups matched the data better than models with isolation and ghost populations. Ragsdale and coworkers call their range of models weakly-structured stem models. In such models, today's populations all share a common ancestry from groups that mixed over time, somewhat separate but connected by gene flow. For these models to fit today's genomes, the separation with gene flow had to persist for a very long time—starting up to a million years ago.
When I studied these models in the paper, what I found most interesting was the branching of recent groups across Africa. Since the 2017 work by Schlebusch and coworkers, most models have included an early differentiation of southern African from other African groups—placing the node that connects these living people as early as 300,000 years ago. The coupling with the age of the Jebel Irhoud fossils has led some anthropologists to think of this as the “origin” of our species. But Ragsdale and coworkers find a much more recent differentiation of today's groups, placing it a bit more than 100,000 years ago.
The reason for this difference in models is fairly intuitive. In a model with high isolation, most of the present-day diversity across Africa has to come from the initial branching of today's groups, the ones represented by genome data. So a lot of ancient between-group variation must mean a deep divergence. Ghost populations can contribute a small amount of variation, but a ghost footprint cannot be too large or it will simply look like a deep divergence between the groups that aren't ghosts. By contrast, a model built on gene flow can have ancient high variation without isolation. In the weakly-structed stem models, the differentiation of today's groups can be a lot more recent, because some ancient variation between them can come from the ancestral stem that they share.
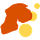
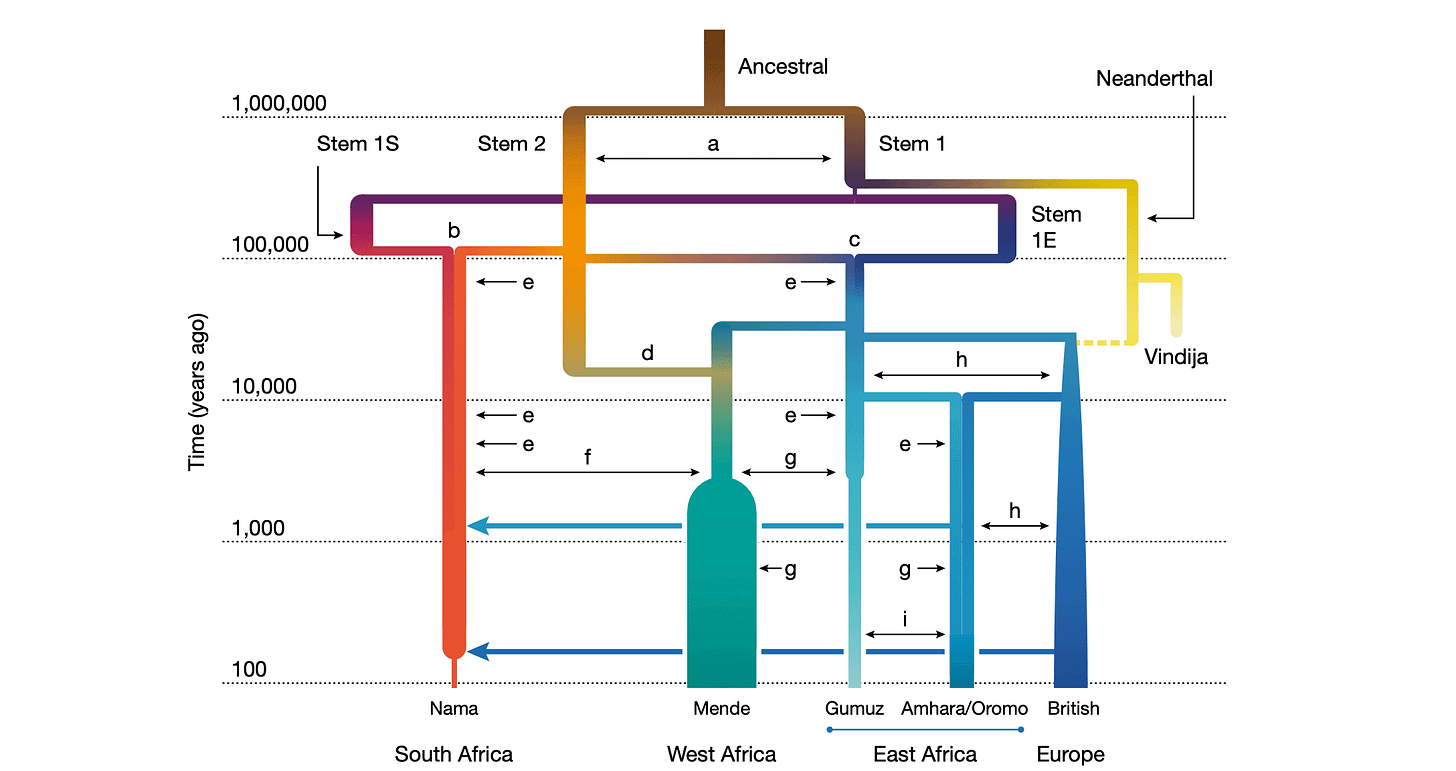
As they tinkered with different kinds of models, Ragsdale and coworkers found a complicated nesting of ancestral groups that fit today's genomes better than any other. In this model, the ancestry of southern African groups like the Nama coalesced from two stem populations around 100,000 years ago. At around the same time, the ancestors of other populations coalesced from one of these stems and yet a third. The branch leading to Neandertals itself emerged from this deep structure, one bottlenecked thread ultimately spilling across Africa into every living group.
When you look at the details of this model, recent events are not so different in their topology from models with ghost populations. For example, in this model the “stem 2” population contributes late to the ancestry of western African people, sometime before 10,000 years ago. This plays a similar role to the “ghost archaics” found in models with more isolation. But in the weakly-structured stem model, this stem population was never isolated from all others. It was one of several that interacted repeatedly during the earlier history of our species.
In other words, this model is truly a braided stream. The groups that form early in our species' evolution continue to play new roles as time passes, remixing with each other in different combinations. These ancestral groups existed long before any fossils with anatomy that paleoanthropologists have recognized within the range of variation of recent people. Our transformation into the form that people around the world share today took place over a million years of interactions.
Many geneticists in the last years of the twentieth century thought that our species underwent a tight bottleneck at some time in the last 200,000 years. Much of the fuel for this idea came from analyses of the mitochondrial DNA and Y chromosome. But as geneticists increased the representation of African populations, they began to find that these populations did not show the same signs of a founder effect as populations of Eurasia. Henry Harpending and collaborators suggested a scenario in which African populations diversified over many thousands of years before a global expansion, which they termed the “Weak Garden of Eden” model. Their 1993 paper was the first expression of a model that combined an African origin with a structured stem population.
Fifteen years later, researchers could examine data from across the nuclear genome, which reinforced that the diversity of African populations had endured for a very long time. More researchers began to think about the relationship of population structure and population growth, trying to find a combination of isolation and demographic growth that might fit the data better. One solution, proposed by Michael Blum and Mattias Jakobsson, was a structured stem population.
“We show that an ancestral bottleneck in Africa, possibly arising in a structured population, can account for the unexpectedly large discrepancy between young mtDNA and Y chromosome ancestors and old autosomal and X-linked ancestors.”—Michael Blum and Mattias Jakobsson (2010)
The weakly-structured stem idea helps broaden today's conversation to include these ways of thinking from more than a decade ago. In some of my own research, published in 2017, I found that the structured stem population and introgression from long-separated groups both were viable explanations for some aspects of genomic variation.
Crossing the steams
While these models differ a lot in the way they assume ancient human groups behaved, the mathematical differences between them are not very great. Both models with ghost populations and structured stem models share the insight that recent populations emerged from the interaction of groups that lived long before 200,000 years ago. Some of them go back at least 700,000 years or longer.
That makes it very important to evaluate these models with other kinds of evidence. Fossil and archaeological material are crucial to understand the succession of ancient populations, and a number of specialists in the Middle Stone Age archaeology of Africa have already begun to use structured metapopulation models in their work. Leaders in this area of research include Eleanor Scerri, who has emphasized that shared material culture across large parts of Africa during the Middle Stone Age may reflect population contacts across the continent.
Readers know that I've long been philosophically aligned with this way of looking at ancient human groups. Gene flow and cultural interactions were part of the landscape that made us human. But the fossil record of this crucial time and place presents some tough challenges. The implication of the weakly-structured stem idea is that the genetic structure of humans today began to form long before any fossil evidence of so-called modern humans existed. But the fossils give very little information about populations of Africa between a million and 200,000 years ago. Only a few have clear geological age estimates, and most regions of Africa have no evidence at all. The fossil group about which we have the most evidence—Homo naledi—may be part of a different picture entirely. That being said, I think there is a lot of promise for building a stronger picture from the fossils. The field has been discovering surprising new fossil evidence for quite a while now, and that's not about to end.
For geneticists, one of the most important areas of research is understanding the connections of Holocene populations within Africa. New studies almost every month are adding more and more data from reseach participants across Africa, representing both large urban populations and small-scale societies. Just in the last couple of months, that includes major studies by Shaohua Fan and coworkers from Sarah Tishkoff's research group, and Nancy Bird and collaborators, both groups including analysis of Holocene interactions. These are important. Scientists who are building population models need to ground those in real data about how human populations have interacted, and those data can only come from more detailed studies of how human population movements and interactions have actually happened.
Notes: This is a fast-developing area and I will be writing more about it in the near future. While I wanted to be sure to mention some of the fossil and archaeological work, this has its own history.
One thing I often observe in reading new work is that today's researchers sometimes neglect earlier authors who wrote about the same problems. Population models have been around for a very long time, after all, and even poor datasets like mtDNA restriction site polymorphisms once prompted many of the same questions people are asking today about whole genomes. I'll continue to try to make some of those connections that are being omitted from recent review articles on structured metapopulation models.
References
Bird, N., Ormond, L., Awah, P., Caldwell, E. F., Connell, B., Elamin, M., Fadlelmola, F. M., Matthew Fomine, F. L., López, S., MacEachern, S., Moñino, Y., Morris, S., Näsänen-Gilmore, P., Nketsia V, N. K., Veeramah, K., Weale, M. E., Zeitlyn, D., Thomas, M. G., Bradman, N., & Hellenthal, G. (2023). Dense sampling of ethnic groups within African countries reveals fine-scale genetic structure and extensive historical admixture. Science Advances, 9(13), eabq2616. https://doi.org/10.1126/sciadv.abq2616
Blum, M. G. B., & Jakobsson, M. (2011). Deep Divergences of Human Gene Trees and Models of Human Origins. Molecular Biology and Evolution, 28(2), 889–898. https://doi.org/10.1093/molbev/msq265
Durvasula, A., & Sankararaman, S. (2020). Recovering signals of ghost archaic introgression in African populations. Science Advances, 6(7), eaax5097. https://doi.org/10.1126/sciadv.aax5097
Fan, S., Spence, J. P., Feng, Y., Hansen, M. E. B., Terhorst, J., Beltrame, M. H., Ranciaro, A., Hirbo, J., Beggs, W., Thomas, N., Nyambo, T., Mpoloka, S. W., Mokone, G. G., Njamnshi, A. K., Fokunang, C., Meskel, D. W., Belay, G., Song, Y. S., & Tishkoff, S. A. (2023). Whole-genome sequencing reveals a complex African population demographic history and signatures of local adaptation. Cell, 186(5), 923-939.e14. https://doi.org/10.1016/j.cell.2023.01.042
Harpending, H. C., Sherry, S. T., Rogers, A. R., & Stoneking, M. (1993). The Genetic Structure of Ancient Human Populations. Current Anthropology, 34(4), 483–496. https://doi.org/10.1086/204195
Hawks, J. (2017). Introgression Makes Waves in Inferred Histories of Effective Population Size. Human Biology, 89(1), 67–80. https://doi.org/10.13110/humanbiology.89.1.04
Lachance, J., Vernot, B., Elbers, C. C., Ferwerda, B., Froment, A., Bodo, J.-M., Lema, G., Fu, W., Nyambo, T. B., Rebbeck, T. R., Zhang, K., Akey, J. M., & Tishkoff, S. A. (2012). Evolutionary History and Adaptation from High-Coverage Whole-Genome Sequences of Diverse African Hunter-Gatherers. Cell, 150(3), 457–469. https://doi.org/10.1016/j.cell.2012.07.009
Lipson, M., Ribot, I., Mallick, S., Rohland, N., Olalde, I., Adamski, N., Broomandkhoshbacht, N., Lawson, A. M., López, S., Oppenheimer, J., Stewardson, K., Asombang, R. N., Bocherens, H., Bradman, N., Culleton, B. J., Cornelissen, E., Crevecoeur, I., de Maret, P., Fomine, F. L. M., … Reich, D. (2020). Ancient West African foragers in the context of African population history. Nature, 577(7792), Article 7792. https://doi.org/10.1038/s41586-020-1929-1
Pfennig, A., Petersen, L. N., Kachambwa, P., & Lachance, J. (2023). Evolutionary genetics and admixture in African populations. Genome Biology and Evolution, evad054. https://doi.org/10.1093/gbe/evad054
Plagnol, V., & Wall, J. D. (2006). Possible Ancestral Structure in Human Populations. PLOS Genetics, 2(7), e105. https://doi.org/10.1371/journal.pgen.0020105
Ragsdale, A. P., Weaver, T. D., Atkinson, E. G., Hoal, E. G., Möller, M., Henn, B. M., & Gravel, S. (2023). A weakly structured stem for human origins in Africa. Nature, 1–9. https://doi.org/10.1038/s41586-023-06055-y
Scerri, E. M. L., Chikhi, L., & Thomas, M. G. (2019). Beyond multiregional and simple out-of-Africa models of human evolution. Nature Ecology & Evolution, 3(10), Article 10. https://doi.org/10.1038/s41559-019-0992-1
Scerri, E. M. L., Thomas, M. G., Manica, A., Gunz, P., Stock, J. T., Stringer, C., Grove, M., Groucutt, H. S., Timmermann, A., Rightmire, G. P., d’Errico, F., Tryon, C. A., Drake, N. A., Brooks, A. S., Dennell, R. W., Durbin, R., Henn, B. M., Lee-Thorp, J., deMenocal, P., … Chikhi, L. (2018). Did Our Species Evolve in Subdivided Populations across Africa, and Why Does It Matter? Trends in Ecology & Evolution, 33(8), 582–594. https://doi.org/10.1016/j.tree.2018.05.005
Schlebusch, C. M., Malmström, H., Günther, T., Sjödin, P., Coutinho, A., Edlund, H., Munters, A. R., Vicente, M., Steyn, M., Soodyall, H., Lombard, M., & Jakobsson, M. (2017). Southern African ancient genomes estimate modern human divergence to 350,000 to 260,000 years ago. Science, 358(6363), 652–655. https://doi.org/10.1126/science.aao6266